Data manipulation is a critical, core skill in data science.
If you want to get a job as a data scientist, you need to master basic data manipulation operations. Ideally, you should be able to write them rapidly, and from memory (no looking them up on Google!).
A very common data manipulation task is manipulating columns of a dataframe. Specifically, you need to know how to add a column to a dataframe.
Adding a column to a dataframe in R is not hard, but there are a few ways to do it. This can make it a little confusing for beginners … you might see several different ways to add a column to a dataframe, and it might not be clear which one you should use.
That being the case, I’m going to show you two very simple techniques to do this, with a specific focus on the method I think is “the best.”
First I’ll show you how to add a column to a dataframe using dplyr. I’ll show you this first, because dplyr is definitely my preferred method. (If you don’t use dplyr, you should … it’s awesome.)
Second, I’ll show you how to add a column to a dataframe with base R. I don’t really like the base R method (it’s a little kludgy and “old fashioned”) but I still want you to see it.
Let’s start with the dplyr method.
Add a column to a dataframe in R using dplyr
In my opinion, the best way to add a column to a dataframe in R is with the mutate() function from dplyr.
mutate(), like all of the functions from dplyr is easy to use.
Let’s take a look:
Load packages
First things first: we’ll load the packages that we will use. Specifically, we’ll load dplyr and caret. dplyr has the mutate() function that we will use, and the caret package has the dataset that we will be working with, the Sacramento dataframe.
#-------------- # LOAD PACKAGES #-------------- library(dplyr) library(caret) library(tidyverse)
Load the dataset
Next, we’ll load our dataset. We’ll be working with the Sacramento dataframe from the caret package.
To load this dataset, we’ll use the data() function.
#-----------------------
# LOAD 'Sacramento' DATA
#-----------------------
data("Sacramento")
Rename the data
Very quickly, before moving on, I’m going to rename the dataset.
This is a minor thing, but little details can make a difference. We’ll rename this for two minor reasons. First, I typically like to avoid capital letters in variable names and dataset names. Second, the name “Sacramento” is not very descriptive. Because of these two reasons, I’ll rename the dataframe to sacramento_housing.
#------------ # RENAME DATA #------------ sacramento_housing <- Sacramento
Inspect the data
Let's also quickly take a look at the data.
sacramento_housing %>% glimpse() # Observations: 932 # Variables: 9 # $ citySACRAMENTO, SACRAMENTO, SACRAMENTO, SACRAMENTO, SACRAMENTO, SACRAMENTO... # $ zip z95838, z95823, z95815, z95815, z95824, z95841, z95842, z95820, z95670... # $ beds 2, 3, 2, 2, 2, 3, 3, 3, 2, 3, 3, 3, 1, 3, 2, 2, 2, 2, 2, 3, 3, 4, 4, 3,... # $ baths 1, 1, 1, 1, 1, 1, 2, 1, 2, 2, 2, 2, 1, 1, 2, 2, 1, 1, 1, 2, 2, 2, 2, 2,... # $ sqft 836, 1167, 796, 852, 797, 1122, 1104, 1177, 941, 1146, 909, 1289, 871, ... # $ type Residential, Residential, Residential, Residential, Residential, Condo... # $ price 59222, 68212, 68880, 69307, 81900, 89921, 90895, 91002, 94905, 98937, 1... # $ latitude 38.63191, 38.47890, 38.61830, 38.61684, 38.51947, 38.66260, 38.68166, 3... # $ longitude -121.4349, -121.4310, -121.4438, -121.4391, -121.4358, -121.3278, -121....
Add a new variable using mutate
Now that we have our dataset, let's add a new variable. Here, we will add a variable called price_per_sqft.
#---------------------------- # ADD VARIABLE USING mutate() #---------------------------- sacramento_housing <- mutate(sacramento_housing, price_per_sqft = price/sqft)
Let's quickly unpack this.
You'll see here that we're using the mutate() function. Inside of mutate(), you'll see that we're referencing the sacramento_housing dataframe.
After we specify the dataframe that we're going to mutate, we specify exactly how we will change it. Here, we are simply creating a new variable called price_per_sqft. price_per_sqft is simply a calculated variable. We are calculating it by dividing the price variable by the sqft variable. Remember, both price and sqft are variables that already exist in the sacramento_housing dataframe.
How the mutate() function works
mutate() is very straightforward to use. Basically, mutate() modifies a dataframe by creating a new variable. That's all that it does.
When you call mutate, the first argument is the name of the dataframe that we want to modify. In the example above, it is the sacramento_housing dataframe.
The second argument is a "name value pair." That sounds a little cryptic, but it's not that complicated. "Name value pair" just means that we're creating a new variable with a "name" and we're assigning some value to that new name. A name and a value. A variable name and a value associated with it. Name value pair.
The variable that we create can be relatively simple or complex. The above example is pretty straightforward. We're just dividing one variable in the dataframe by another. That's the "value" that we're calculating, and we're giving it the name price_per_sqft.
However, new variables can be rather complicated as well. We can use a variety of mathematical functions and logical functions to calculate the value of the new variable. I won't go into that right now, but understand that you have a lot of flexibility concerning how you calculate the values of the new variables you create.
Add a column to a dataframe in R using "base R"
Now, I'll show you a way to add a new column to a dataframe using base R.
Before we get into it, I want to make a few comments.
First, there are several different ways to add a new variable to a dataframe using base R. I'll show you only one.
Second, using base R to add a new column to a dataframe is not my preferred method. I strongly prefer using mutate() from dplyr (I'll discuss why I prefer dplyr below).
With those comments in mind, let's walk through how to add a new column to a dataframe using base R.
Create a new dataframe
First, we will create a new dataframe using the tribble() function.
After creating it, we'll quickly print out the data just to inspect it.
#========================================== # ADD A COLUMN TO A DATAFRAME USING BASE R # - this is data about city-level nominal # GDP taken from Wikipedia # - Note that these quanities are estimated # nominal GDP for either 2015 or 2016 #========================================== # source: https://en.wikipedia.org/wiki/List_of_cities_by_GDP top_5_city_gdp <- tribble( ~city, ~gdp_billion_dollars, 'Tokyo',1893 ,'New York',1657 ,'Los Angeles',1001 ,'Paris',850 ,'Chicago',651 ) # INSPECT top_5_city_gdp # A tibble: 5 x 2 # city gdp_billion_dollars ## 1 Tokyo 1893 # 2 New York 1657 # 3 Los Angeles 1001 # 4 Paris 850 # 5 Chicago 651
You can see that the dataframe only has two variables: city and gdp_billion_dollars.
Add a new column to the dataframe
Now, we'll add a new column to the dataframe. The new variable will be called country, and it will simply contain the name of the country.
To do this, we're going to use the '$' operator. This normally allows us to reference the name of a column in a dataframe. When we use the $ operator, we specify the dataframe first, then the $ symbol, then the name of the variable. So using this operator takes the form:
dataframe$variable
However, in this case, we can actually use it to create a new variable.
Let's take a look:
#============================
# ADD NEW COLUMN TO DATAFRAME
#============================
top_5_city_gdp$country <- c('Japan'
,'United States'
,'United States'
,'France'
,'United States'
)
# INSPECT
top_5_city_gdp
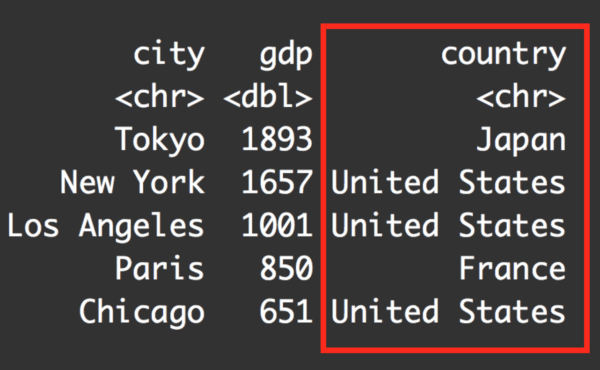
# # A tibble: 5 x 3
# city gdp_billion_dollars country
#
# 1 Tokyo 1893 Japan
# 2 New York 1657 United States
# 3 Los Angeles 1001 United States
# 4 Paris 850 France
# 5 Chicago 651 United States
Notice that the dataframe now has the new variable, country.
So how does this work?
The code top_5_city_gdp$country basically creates a new variable, country, and we're assigning the values using the assignment operator, <-. The new values are contained within a vector that we have created using the c() function.
Why you should use the Tidyverse for data science in R
As I mentioned earlier, I strongly prefer using mutate() to add a column to a dataframe in R. In fact, for most data manipulation tasks and data science tasks, I think the functions from dplyr and the Tidyverse are superior.
If you're not familiar, the "Tidyverse" is a set of packages for doing data science and data analysis in the R programming language.
The Tidyverse packages include:
dplyrfor data manipulationggplot2for data visualizationtidyrfor data reshaping (which is a sub-task of data manipulation)lubridateto deal with datesforcatsto deal with R's factor variable typestringrto deal with string data
There are also a few other packages in the Tidyverse, but these are the core.
Why dplyr and the Tidyverse are better than many other R tools
The reason that I prefer the tools from the Tidyverse packages (like using mutate() to add new variables) is that they are easy to use.
Almost all of the functions from dplyr and the Tidyverse read like pseudocode. When you want to add a variable to a dataframe, you "mutate" it by using the mutate() function. When you want to subset your data, you "filter" it by using the filter() function. Nearly all of the functions in dplyr and the Tidyverse are very well named. This makes them easy to learn, easy to remember, and easy to use.
Moreover, the functions of the Tidyverse do one thing and one thing only. For example, mutate() only does one thing: it adds new variables to a dataframe. Because the Tidyverse functions only do one thing, you can use them almost like LEGO building blocks. Once you start using the Tidyverse, you realize how well designed it is.
This is why R's "Tidyverse" packages are great.
If you're getting started with data science in R, I strongly recommend that you focus on learning the Tidyverse. That means you should learn ggplot2, dplyr, stringr, tidyr, forcats, and a few others.
For better or worse, there are many different way to accomplish data science tasks in R.
I'll get hate mail for saying this, but I strongly think that the tools of the Tidyverse are better than the base R methods or other methods. You'll save yourself a lot of time and frustration by learning the Tidyverse way of doing things verses the older methodologies from R.
That's not to say that you can completely ignore base R. You will still need to know some tools and methods from base R. However, if there's a choice between doing something with a Tidyverse tool (like dplyr::mutate()) or base R, I strongly suggest that you use the Tidyverse method. Again, the tools of the Tidyverse are easier to read, easier to learn, easier to use, and easier to debug.
Learn and master the core tools of data science in R
Here at Sharp Sight, we teach data science.
We'll teach you how to do data science in R by using tools like dplyr, mutate(), and the other data science tools of R's Tidyverse.
If you're ready to learn and master data science in R, sign up for our email list.
If you sign up, you'll get free data science tutorials, delivered every week to your inbox.
You'll also hear about our premium data science courses when they open for enrollment.
If you're serious about learning and mastering data science as fast as possible, sign up now.
I totally agree with you, that anyone who is currently learning R should consider and learn the newer approaches and strategies as included in the Tidyverse. Even so anybody who learned R long time ago when it wasn’t present should keep its eyes open to check if there is something better around than base R.
However for the sake of the argument it would be nice to show the same example with both ways. In fact adding a vector as column is easier with base R and performing column-wise calculations easier with dplyr.
I took some time to learn tidyverse but noticed that other important functions that I had written would not work with the tidyverse-altered data. With that being said, once you’re in, you’re in. Tidyverse may or may not be able to do some of the things you want it to do and writing base-R functions to get around that may not be an option.
I was working with categorical data, for the record, which may be a large part of the reason for my issues.
Were you using forcats and stringers to manipulate your factor/categorical variables?
Stupid question time… I am using dplyr and mutate to create a new column in my dataset, but when I then print the dataset, it’s not there. Here’s my code:
Is this what I should expect? How do I make the new column stick around?
The source of this issue is really about the output of
mutate().mutate()does not directly modify the original dataframe (i.e.,my_register).Instead,
mutate()produces a new dataframe that contains the new column. By default, this new dataframe is sent to the console, which means that it’s printed out in the console but not saved.If you want to save the output, you need to use an assignment operation to store the output to a name (i.e.,
<-).That's the common way to do it. But the Tidyverse also has another assignment operator that you can use at the end of a dplyr chain. You can use the
->operator like this:I prefer this second version because it's easier to read from top to bottom.
Keep in mind that in both examples, I've used the name
my_register_UPDATEDso that I'm not overwriting the original dataset.To be clear: you can overwrite the original, but you need to be careful. Always test your code to make sure that it's working correctly before you overwrite your data.
When you have it tested and working properly, you can use the following to store the output of the operation with the name of the original dataset.